R plugin (Preview)
The R plugin runs a user-defined function (UDF) using an R script.
The script gets tabular data as its input, and produces tabular output. The plugin's runtime is hosted in a sandbox on the cluster's nodes. The sandbox provides an isolated and secure environment.
Syntax
T | evaluate [hint.distribution = (single | per_node)] r(output_schema, script [, script_parameters] [, external_artifacts])
Learn more about syntax conventions.
Parameters
| Name | Type | Required | Description |
|---|---|---|---|
| output_schema | string |
✔️ | A type literal that defines the output schema of the tabular data, returned by the R code. The format is: typeof(ColumnName: ColumnType[, ...]). For example: typeof(col1:string, col2:long). To extend the input schema, use the following syntax: typeof(*, col1:string, col2:long). |
| script | string |
✔️ | The valid R script to be executed. |
| script_parameters | dynamic |
A property bag of name and value pairs to be passed to the R script as the reserved kargs dictionary. For more information, see Reserved R variables. |
|
hint.distribution |
string |
Hint for the plugin's execution to be distributed across multiple cluster nodes. The default value is single. single means that a single instance of the script will run over the entire query data. per_node means that if the query before the R block is distributed, an instance of the script will run on each node over the data that it contains. |
|
| external_artifacts | dynamic |
A property bag of name and URL pairs for artifacts that are accessible from cloud storage. They can be made available for the script to use at runtime. URLs referenced in this property bag are required to be included in the cluster's callout policy and in a publicly available location, or contain the necessary credentials, as explained in storage connection strings. The artifacts are made available for the script to consume from a local temporary directory, .\Temp. The names provided in the property bag are used as the local file names. See Example. For more information, see Install packages for the R plugin. |
Reserved R variables
The following variables are reserved for interaction between Kusto Query Language and the R code:
df: The input tabular data (the values ofTabove), as an R DataFrame.kargs: The value of the script_parameters argument, as an R dictionary.result: An R DataFrame created by the R script. The value becomes the tabular data that gets sent to any Kusto query operator that follows the plugin.
Enable the plugin
- The plugin is disabled by default.
- Enable or disable the plugin in the Azure portal in the Configuration tab of your cluster. For more information, see Manage language extensions in your Azure Data Explorer cluster (Preview)
R sandbox image
- The R sandbox image is based on R 3.4.4 for Windows, and includes packages from Anaconda's R Essentials bundle.
Examples
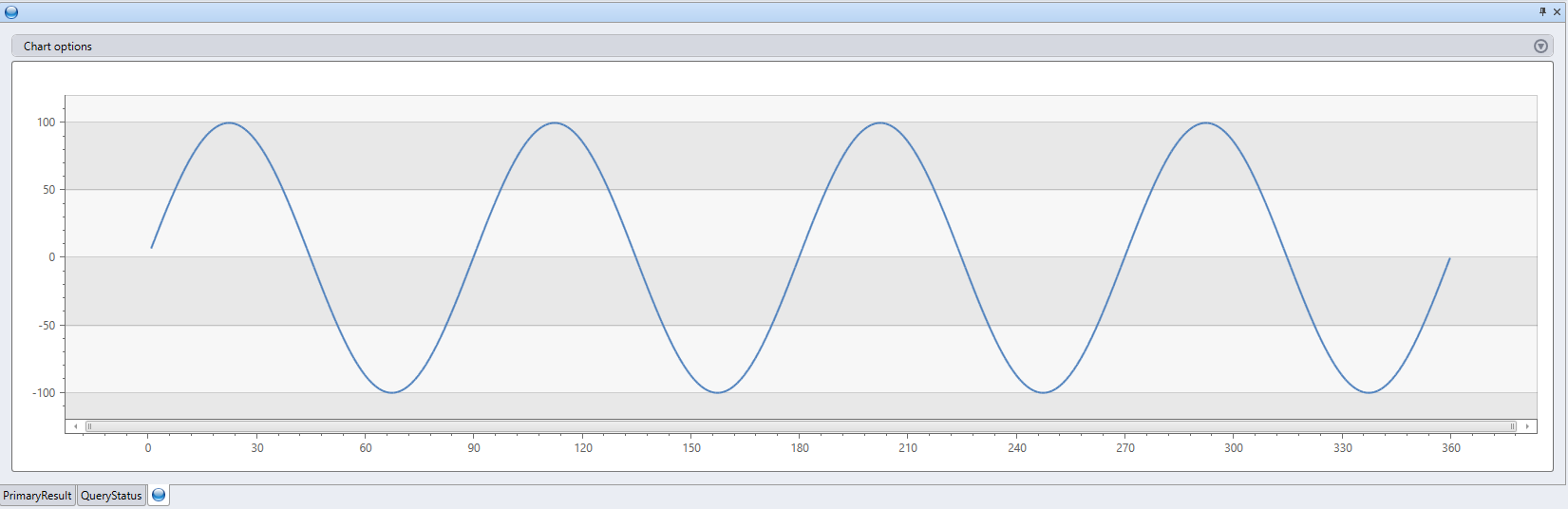
range x from 1 to 360 step 1
| evaluate r(
//
typeof(*, fx:double), // Output schema: append a new fx column to original table
//
'result <- df\n' // The R decorated script
'n <- nrow(df)\n'
'g <- kargs$gain\n'
'f <- kargs$cycles\n'
'result$fx <- g * sin(df$x / n * 2 * pi * f)'
//
, bag_pack('gain', 100, 'cycles', 4) // dictionary of parameters
)
| render linechart
Performance tips
Reduce the plugin's input dataset to the minimum amount required (columns/rows).
Use filters on the source dataset using the Kusto Query Language, when possible.
To make a calculation on a subset of the source columns, project only those columns before invoking the plugin.
Use
hint.distribution = per_nodewhenever the logic in your script is distributable.You can also use the partition operator for partitioning the input data et.
Whenever possible, use the Kusto Query Language to implement the logic of your R script.
For example:
.show operations | where StartedOn > ago(1d) // Filtering out irrelevant records before invoking the plugin | project d_seconds = Duration / 1s // Projecting only a subset of the necessary columns | evaluate hint.distribution = per_node r( // Using per_node distribution, as the script's logic allows it typeof(*, d2:double), 'result <- df\n' 'result$d2 <- df$d_seconds\n' // Negative example: this logic should have been written using Kusto's query language ) | summarize avg = avg(d2)
Usage tips
To avoid conflicts between Kusto string delimiters and R string delimiters:
- Use single quote characters (
') for Kusto string literals in Kusto queries. - Use double quote characters (
") for R string literals in R scripts.
- Use single quote characters (
Use the external data operator to obtain the content of a script that you've stored in an external location, such as Azure blob storage or a public GitHub repository.
For example:
let script = externaldata(script:string) [h'https://kustoscriptsamples.blob.core.windows.net/samples/R/sample_script.r'] with(format = raw); range x from 1 to 360 step 1 | evaluate r( typeof(*, fx:double), toscalar(script), bag_pack('gain', 100, 'cycles', 4)) | render linechart
Install packages for the R plugin
Follow these step by step instructions to install package(s) that aren't included in the plugin's base image.
Prerequisites
Create a blob container to host the packages, preferably in the same place as your cluster. For example,
https://artifactswestus.blob.core.windows.net/r, assuming your cluster is in West US.Alter the cluster's callout policy to allow access to that location.
This change requires AllDatabasesAdmin permissions.
For example, to enable access to a blob located in
https://artifactswestus.blob.core.windows.net/r, run the following command:
.alter-merge cluster policy callout @'[ { "CalloutType": "sandbox_artifacts", "CalloutUriRegex": "artifactswestus\\.blob\\.core\\.windows\\.net/r/","CanCall": true } ]'
Install packages
The example snips below assume local R machine on Windows environment.
Verify you're using the appropriate R version – current R Sandbox version is 3.4.4:
> R.Version()["version.string"] $version.string [1] "R version 3.4.4 (2018-03-15)"If needed you can download it from here.
Launch the x64 RGui
Create a new empty folder to be populated with all the relevant packages you would like to install. In this example we install the brglm2 package, so creating "C:\brglm2".
Add the newly created folder path to lib paths:
> .libPaths("C://brglm2")Verify that the new folder is now the first path in .libPaths():
> .libPaths() [1] "C:/brglm2" "C:/Program Files/R/R-3.4.4/library"Once this setup is done, any package that we install shall be added to this new folder. Let's install the requested package and its dependencies:
> install.packages("brglm2")In case the question "Do you want to install from sources the packages which need compilation?" pops up, answer "Y".
Verify that new folders were added to "C:\brglm2":
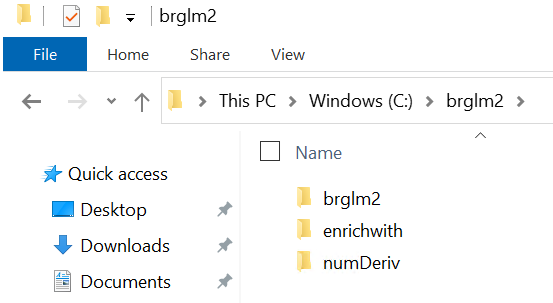
Select all items in that folder and zip them to e.g. libs.zip (do not zip the parent folder). You should get an archive structure like this:
libs.zip:
- brglm2 (folder)
- enrichwith (folder)
- numDeriv (folder)
Upload libs.zip to the blob container that was set above
Call the
rplugin.- Specify the
external_artifactsparameter with a property bag of name and reference to the zip file (the blob's URL, including a SAS token). - In your inline r code, import
zipfilefromsandboxutilsand call itsinstall()method with the name of the zip file.
- Specify the
Example
Install the brglm2 package:
print x=1
| evaluate r(typeof(*, ver:string),
'library(sandboxutils)\n'
'zipfile.install("brglm2.zip")\n'
'library("brglm2")\n'
'result <- df\n'
'result$ver <-packageVersion("brglm2")\n'
,external_artifacts=bag_pack(brglm2.zip', 'https://artifactswestus.blob.core.windows.net/r/libs.zip?*** REPLACE WITH YOUR SAS TOKEN ***'))
| x | ver |
|---|---|
| 1 | 1.8.2 |
Make sure that the archive's name (first value in pack pair) has the *.zip suffix to prevent collisions when unzipping folders whose name is identical to the archive name.
This capability isn't supported.
Feedback
Coming soon: Throughout 2024 we will be phasing out GitHub Issues as the feedback mechanism for content and replacing it with a new feedback system. For more information see: https://aka.ms/ContentUserFeedback.
Submit and view feedback for